Qingchu Jin, PhD
Faculty Scientist I
Center for Molecular Medicine
Jin Lab
Developing trustworthy artificial intelligence (AI) systems and computational approaches to improve patient care and to uncover the biological mechanisms that shape human health.
The past decade has seen extraordinary advances in AI, driven by rapid progress in computing infrastructure. At the same time, medicine and biology have entered an era of enormous data availability. Hospitals now collect massive, high-resolution clinical data streams, while modern biological research generates large-scale multi-omics datasets. These developments create remarkable opportunities. Yet they also raise fundamental challenges. In healthcare, AI systems operate in high-stakes environments where uncertainty matters. In biology, high-dimensional multi-omics data require models that can not only make prediction but also phenotype the samples, identify the trajectory, and uncover the mechanism.
The Jin lab works at the intersection of these domains. We build computational frameworks that are not only accurate, but safe, interpretable, and biologically meaningful.
AI for Healthcare
Clinical data are complex, multimodal, and dynamic. Electronic health records, physiological waveforms, laboratory measurements, and imaging data together form a longitudinal portrait of patient health. We develop multimodal, real-time machine learning (ML) models that integrate these diverse data streams to assess patient risk and monitor disease progression.
A central focus of our lab is safe AI. Even highly accurate models are imperfect, and in medicine, a single incorrect decision can have serious consequences. We develop computational approaches that incorporate uncertainty quantification and confidence estimation at the individual patient level. By explicitly modeling predictive uncertainty, we aim to facilitate safer clinical deployment and responsible implementation.
Beyond predicting the patients’ outcome, our lab also is interested in causal ML models that estimate the potential effect of treatment strategies. Unlike traditional predictive models that forecast outcomes, causal models aim to support decision-making by evaluating what may happen under different interventions. This moves AI closer to personalized treatment planning and real-world clinical decision.
Our work is deeply embedded within the healthcare system through close collaboration with MaineHealth Maine Medical Center: 1) In the cardiovascular ICU, we are developing multimodal real-time models to monitor recovery following cardiac surgery (Figure 1). 2) In the medical ICU, we are building causal machine learning models to support treatment strategies for severe hyponatremia (Figure 2).
Through these translational partnerships, our lab seeks to bridge the gap between algorithm development and bedside implementation.
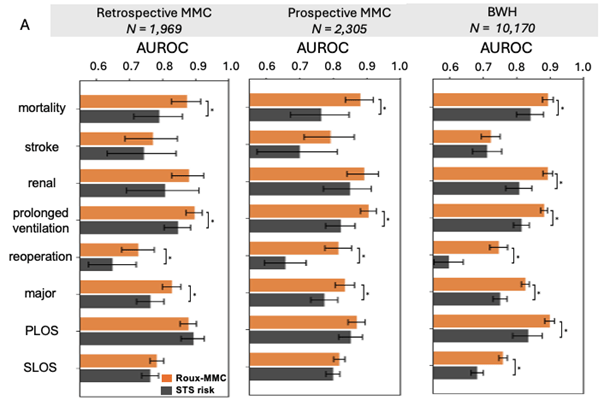
Figure 1
Performance comparison between the Roux–MMC multimodal model and the Society of Thoracic Surgeons (STS) risk calculator for predicting postoperative outcomes in cardiac surgery patients. The Roux–MMC model incorporates both preoperative and intraoperative data, whereas the STS calculator is based solely on preoperative variables. Model discrimination is evaluated using the area under the receiver operating characteristic curve (AUROC).
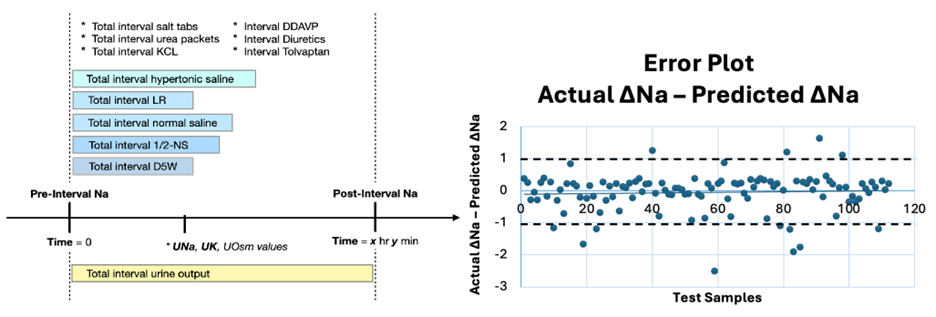
Figure 2
Predictive modeling of treatment effects in patients with severe hyponatremia. The left panel illustrates the longitudinal structure of the data, including sodium measurements before and after treatment initiation. The right panel presents the performance of our hybrid model, an ensemble model combining AI model and mechanistic model, comparing the predicted change in serum sodium concentration with the observed change following treatment.
Computational Biology
We are interested in understanding developmental trajectories using early-life multi-omics data. One key research direction in our lab is modeling childhood obesity growth trajectories using early-life multi-omics signatures. We are specifically interested in the following. How to robustly phenotyping different growth trajectory patterns? Is there any early-life multi-omics signature that can predict the future trajectory pattern and what is the mechanism? To comprehensively study this project, we obtained a wide spectrum omics data from a longitudinal cohort including genomics, methylation, transcriptomics and lipidomics. We believe early accurate prediction of growth pattern can lead to personalized proactive lifestyle and pharmacological intervention to avoid obesity. Figure 3 shows preliminary results for predictive results on mice data. This work is conducted in collaboration with A*STAR in Singapore, Roux Institute at Northeastern University and Van Andel Institute using the GUSTO birth cohort dataset.
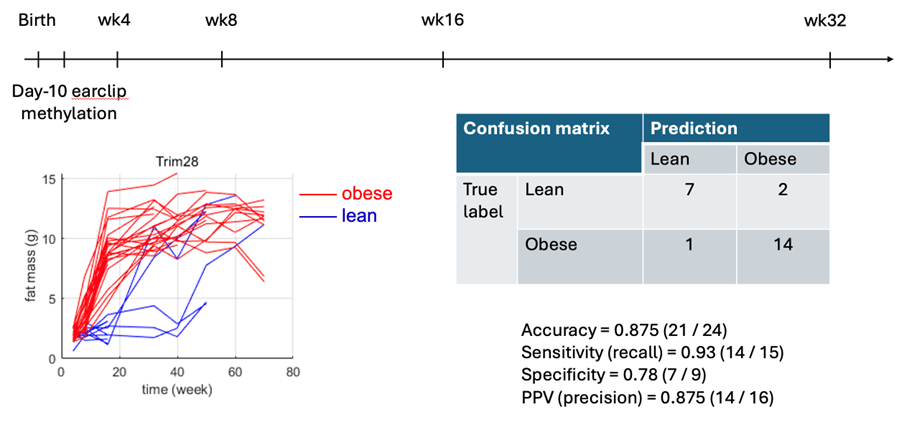
Figure 3
Preliminary longitudinal analysis of 24 Trim28+/D9 heterozygous mutant mice. Obese and lean phenotypes were defined at week 16 based on lean and fat mass measurements (lower left panel). Day-10 DNA methylation array data were used to predict later metabolic phenotype. Using a leave-one-out cross-validation machine learning framework, we applied the first principal component derived from principal component analysis (PCA) as the predictive feature and correctly classified 21 of 24 mice. Detailed performance metrics are shown in the lower right panel.
The Jin Lab is currently actively recruiting a Postdoctoral Fellow and a Master-level Data Analyst with experience in AI in healthcare or multi-omics data analysis. If you are interested, please email qingchu.jin@mainehealth.org and cc qjin.jobapp@gmail.com with your CV and a brief description of your research experience. Please include the job title you are applying for (Postdoctoral Fellow or Data Analyst) in the email title.